Engineering Students Join the Quest to Save Lives
Top row: Jacob Durant and Daniel Wieland. Middle row: Megan Johnson and Anna Peckham. Bottom row: Khaled Albalol, and Jacob Padilla
Project Title: Developing Diagnostics for Monitoring COVID-19 Patients on Ventilators for Secondary Infection
Team 21015 Members:
Khaled Albaloul, industrial engineering
Jacob Thomas Durant, biosystems engineering
Megan Ann Johnson, biomedical engineering
Jacob Celso Padilla, electrical and computer engineering
Anna Marjorie Peckham, mechanical engineering
Daniel Robert Wieland, biomedical engineering, biochemistry, molecular and cellular biology
Sponsor: UA Department of Biosystems Engineering
A study in Clinical Infectious Diseases found that the mortality rate for hospitalized COVID-19 patients in the United States was 21.4% overall, but 70.5% for those on mechanical ventilation. Team 21015 jumped at the chance to help address this vexing issue. For their capstone project, the team is developing a user-friendly application to speed the diagnosis and targeted treatment of secondary infections, such as pneumonia, which are frequently the cause of death for patients on ventilators
The app will display results from a new technology and diagnostic tool called metagenomic next generation sequencing. An evolution of traditional genetic sequencing, this process can identify microbes in the saliva of a patient, including viruses, bacteria, and fungi.
Physicians typically rely on culture testing to target the cause of infection – a process that can take several days. Standard practice is to prescribe a broad-spectrum antibiotic while awaiting results, but that is a non-specific treatment that may lead to antibiotic resistance.
Metagenomic next generation sequencing, however, produces test results in a matter of hours, rather than days.
Diagnosing Secondary Infections Quickly: A Matter of Life and Death
“For patients on ventilators, including many COVID-19 patients, the ability to quickly identify infections and treat them could mean the difference between life and death,” said Megan Johnson, biomedical engineering senior and team lead.
Students are soliciting input from physicians as they design the app. Ideally, they envision a method by which, as soon as the results are ready, a computer or smartphone will display the amount and type of each microbe present. The physician can use this information to promptly prescribe the most effective treatment.
Daniel Wieland, a triple major in biomedical engineering, biochemistry and molecular and cellular biology, is learning RShiny, a data visualization tool the team is using to construct the application. For Wieland, this project is an important building block to his future.
“As a pre-med student with a heavy emphasis on research, this is the type of project I hope to lead as a future physician, for the benefit of my peers and patients alike,” he said.
Skills, Teamwork and a Commitment to Helping
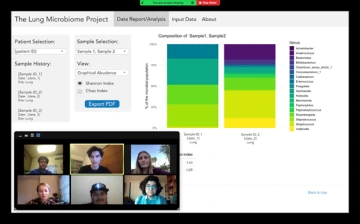
Team 21025 reviews a prototype design for the screen display of their system.
Team members are mastering application software, building and administering surveys, interviewing medical professionals, demonstrating the application interface, and collecting feedback on what viewing options work best. They are also learning about time management, teamwork, and how to accomplish their goal using their individual backgrounds and strengths.
This kind of teamwork is no easy task in ordinary times, and especially difficult during a once-in-a-century pandemic when remote meetings are the only safe option.
“It is amazing to me that we’re working so well together when we have not once met in person,” Johnson said.
Several students commented that it was gratifying to work on a potentially lifesaving project during this tumultuous time of COVID-19.
Jacob Durant, also a biomedical engineering senior, sums it up this way: “Doing this project will be my contribution to the pandemic as an engineer.”

